Receptor tyrosine kinase
| receptor protein-tyrosine kinase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 | |||||||||
| Identifiers | |||||||||
| EC number | 2.7.10.1 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / EGO | ||||||||
| |||||||||
| Identifiers | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Symbol | Pkinase_Tyr | ||||||||
| Pfam | PF07714 | ||||||||
| OPM superfamily | 207 | ||||||||
| OPM protein | 2k1k | ||||||||
| |||||||||
Receptor tyrosine kinases (RTKs) are the high-affinity cell surface receptors for many polypeptide growth factors, cytokines, and hormones. Of the 90 unique tyrosine kinase genes identified in the human genome, 58 encode receptor tyrosine kinase proteins.[1] Receptor tyrosine kinases have been shown not only to be key regulators of normal cellular processes but also to have a critical role in the development and progression of many types of cancer.[2] Receptor tyrosine kinases are part of the larger family of protein tyrosine kinases, encompassing the receptor tyrosine kinase proteins which contain a transmembrane domain, as well as the non receptor tyrosine kinases which do not possess transmembrane domains.[3]
Nomenclature
Receptor tyrosine kinase (RTK) in this article is also known as tyrosine receptor kinase (TRK) or tyrosine kinase receptor (TKR) depending on the permutation.
Receptor tyrosine kinase classes
Approximately 20 different RTK classes have been identified.[4]
- RTK class I (EGF receptor family) (ErbB family)
- RTK class II (Insulin receptor family)
- RTK class III (PDGF receptor family)
- RTK class IV (FGF receptor family)
- RTK class V (VEGF receptors family)
- RTK class VI (HGF receptor family)
- RTK class VII (Trk receptor family)
- RTK class VIII (Eph receptor family)
- RTK class IX (AXL receptor family)
- RTK class X (LTK receptor family)
- RTK class XI (TIE receptor family)
- RTK class XII (ROR receptor family)
- RTK class XIII (DDR receptor family)
- RTK class XIV (RET receptor family)
- RTK class XV (KLG receptor family)
- RTK class XVI (RYK receptor family)
- RTK class XVII (MuSK receptor family)
Structure
Most RTKs are single subunit receptors but some exist as multimeric complexes, e.g., the insulin receptor that forms disulfide linked dimers in the presence of hormone (insulin); moreover, ligand binding to the extracellular domain induces formation of receptor dimers.[5] Each monomer has a single hydrophobic transmembrane-spanning domain composed of 25 to 38 amino acids, an extracellular N terminal region, and an intracellular C terminal region.[6] The extracellular N terminal region exhibits a variety of conserved elements including immunoglobulin (Ig)-like or epidermal growth factor (EGF)-like domains, fibronectin type III repeats, or cysteine-rich regions that are characteristic for each subfamily of RTKs; these domains contain primarily a ligand-binding site, which binds extracellular ligands, e.g., a particular growth factor or hormone.[2] The intracellular C terminal region displays the highest level of conservation and comprises catalytic domains responsible for the kinase activity of these receptors, which catalyses receptor autophosphorylation and tyrosine phosphorylation of RTK substrates.[2]
Kinase activity
In biochemistry, a kinase is a type of enzyme that transfers phosphate groups (see below) from high-energy donor molecules, such as ATP (see below) to specific target molecules (substrates); the process is termed phosphorylation. The opposite, an enzyme that removes phosphate groups from targets, is known as a phosphatase. Kinase enzymes that specifically phosphorylate tyrosine amino acids are termed tyrosine kinases.
When a growth factor binds to the extracellular domain of an RTK, its dimerization is triggered with other adjacent RTKs. Dimerization leads to a rapid activation of the protein's cytoplasmic kinase domains, the first substrate for these domains being the receptor itself. The activated receptor as a result then becomes autophosphorylated on multiple specific intracellular tyrosine residues.
Signal transduction
Through diverse means, extracellular ligand binding will typically cause or stabilize receptor dimerization. This allows a tyrosine in the cytoplasmic portion of each receptor monomer to be trans-phosphorylated by its partner receptor, propagating a signal through the plasma membrane.[7] The phosphorylation of specific tyrosine residues within the activated receptor creates binding sites for Src homology 2 (SH2) domain- and phosphotyrosine binding (PTB) domain-containing proteins.[8][9] Specific proteins containing these domains include Src and phospholipase Cγ. Phosphorylation and activation of these two proteins on receptor binding lead to the initiation of signal transduction pathways. Other proteins that interact with the activated receptor act as adaptor proteins and have no intrinsic enzymatic activity of their own. These adaptor proteins link RTK activation to downstream signal transduction pathways, such as the MAP kinase signalling cascade.[2] An example of a vital signal transduction pathway involves the tyrosine kinase receptor, c-met, which is required for the survival and proliferation of migrating myoblasts during myogenesis. A lack of c-met disrupts secondary myogenesis and—as in LBX1—prevents the formation of limb musculature. This local action of FGFs (Fibroblast Growth Factors) with their RTK receptors is classified as paracrine signalling.
Families
Epidermal growth factor receptor family
The ErbB protein family or epidermal growth factor receptor (EGFR) family is a family of four structurally related receptor tyrosine kinases. Insufficient ErbB signaling in humans is associated with the development of neurodegenerative diseases, such as multiple sclerosis and Alzheimer's Disease.[10] In mice, loss of signaling by any member of the ErbB family results in embryonic lethality with defects in organs including the lungs, skin, heart, and brain. Excessive ErbB signaling is associated with the development of a wide variety of types of solid tumor. ErbB-1 and ErbB-2 are found in many human cancers and their excessive signaling may be critical factors in the development and malignancy of these tumors.[11]
Fibroblast growth factor receptor (FGFR) family
Fibroblast growth factors comprise the largest family of growth factor ligands at 23 members.[12] The natural alternate splicing of four fibroblast growth factor receptor (FGFR) genes results in the production of over 48 different isoforms of FGFR.[13] These isoforms vary in their ligand binding properties and kinase domains; however, all share a common extracellular region composed of three immunoglobulin (Ig)-like domains (D1-D3), and thus belong to the immunoglobulin superfamily.[14] Interactions with FGFs occur via FGFR domains D2 and D3. Each receptor can be activated by several FGFs. In many cases, the FGFs themselves can also activate more than one receptor. This is not the case with FGF-7, however, which can activate only FGFR2b.[13] A gene for a fifth FGFR protein, FGFR5, has also been identified. In contrast to FGFRs 1-4, it lacks a cytoplasmic tyrosine kinase domain, and one isoform, FGFR5γ, only contains the extracellular domains D1 and D2.[15]
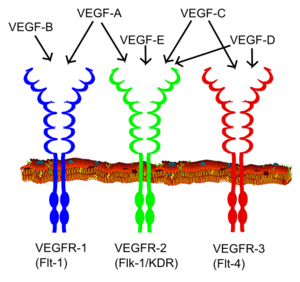
Vascular endothelial growth factor receptor (VEGFR) family
Vascular endothelial growth factor (VEGF) is one of the main inducers of endothelial cell proliferation and permeability of blood vessels. Two RTKs bind to VEGF at the cell surface, VEGFR-1 (Flt-1) and VEGFR-2 (KDR/Flk-1).[16]
The VEGF receptors have an extracellular portion consisting of seven Ig-like domains so, like FGFRs, belong to the immunoglobulin superfamily. They also possess a single transmembrane spanning region and an intracellular portion containing a split tyrosine-kinase domain. VEGF-A binds to VEGFR-1 (Flt-1) and VEGFR-2 (KDR/Flk-1). VEGFR-2 appears to mediate almost all of the known cellular responses to VEGF. The function of VEGFR-1 is less well defined, although it is thought to modulate VEGFR-2 signaling. Another function of VEGFR-1 may be to act as a dummy/decoy receptor, sequestering VEGF from VEGFR-2 binding (this appears to be particularly important during vasculogenesis in the embryo). A third receptor has been discovered (VEGFR-3); however, VEGF-A is not a ligand for this receptor. VEGFR-3 mediates lymphangiogenesis in response to VEGF-C and VEGF-D.
RET receptor family
The natural alternate splicing of the RET gene results in the production of 3 different isoforms of the protein RET. RET51, RET43, and RET9 contain 51, 43, and 9 amino acids in their C-terminal tail, respectively.[17] The biological roles of isoforms RET51 and RET9 are the most well studied in-vivo, as these are the most common isoforms in which RET occurs.
RET is the receptor for members of the glial cell line-derived neurotrophic factor (GDNF) family of extracellular signalling molecules or ligands (GFLs).[18]
In order to activate RET, first GFLs must form a complex with a glycosylphosphatidylinositol (GPI)-anchored co-receptor. The co-receptors themselves are classified as members of the GDNF receptor-α (GFRα) protein family. Different members of the GFRα family (GFRα1-GFRα4) exhibit a specific binding activity for a specific GFLs.[19] Upon GFL-GFRα complex formation, the complex then brings together two molecules of RET, triggering trans-autophosphorylation of specific tyrosine residues within the tyrosine kinase domain of each RET molecule. Phosphorylation of these tyrosines then initiates intracellular signal transduction processes.[20]
Eph receptor family
Ephrin and Eph receptors are the largest subfamily of RTKs.
Discoidin domain receptor (DDR) family
The DDRs are unique RTKs in that they bind to collagens rather than soluble growth factors.[21]
See also
References
- ↑ Robinson DR, Wu YM, Lin SF (November 2000). "The protein tyrosine kinase family of the human genome". Oncogene. 19 (49): 5548–57. doi:10.1038/sj.onc.1203957. PMID 11114734.
- 1 2 3 4 Zwick E, Bange J, Ullrich A (September 2001). "Receptor tyrosine kinase signalling as a target for cancer intervention strategies". Endocrine-Related Cancer. 8 (3): 161–73. doi:10.1677/erc.0.0080161. PMID 11566607.
- ↑ Hubbard SR, Till JH (2000). "Protein tyrosine kinase structure and function". Annual Review of Biochemistry. 69: 373–98. doi:10.1146/annurev.biochem.69.1.373. PMID 10966463.
- ↑ http://www.genome.ad.jp Retrieved on 2007-04-05
- ↑ Lodish; et al. (2003). Molecular cell biology (5th ed.).
- ↑ Hubbard SR (1999). "Structural analysis of receptor tyrosine kinases". Progress in Biophysics and Molecular Biology. 71 (3-4): 343–58. doi:10.1016/S0079-6107(98)00047-9. PMID 10354703.
- ↑ Lemmon MA, Schlessinger J (June 2010). "Cell signaling by receptor tyrosine kinases". Cell. 141 (7): 1117–34. doi:10.1016/j.cell.2010.06.011. PMC 2914105
 . PMID 20602996.
. PMID 20602996. - ↑ Pawson T (February 1995). "Protein modules and signalling networks". Nature. 373 (6515): 573–80. Bibcode:1995Natur.373..573P. doi:10.1038/373573a0. PMID 7531822.
- ↑ Ren S, Yang G, He Y, Wang Y, Li Y, Chen Z (2008). "The conservation pattern of short linear motifs is highly correlated with the function of interacting protein domains". BMC Genomics. 9: 452. doi:10.1186/1471-2164-9-452. PMC 2576256
 . PMID 18828911.
. PMID 18828911. - ↑ Bublil EM, Yarden Y (April 2007). "The EGF receptor family: spearheading a merger of signaling and therapeutics". Current Opinion in Cell Biology. 19 (2): 124–34. doi:10.1016/j.ceb.2007.02.008. PMID 17314037.
- ↑ Cho HS, Leahy DJ (August 2002). "Structure of the extracellular region of HER3 reveals an interdomain tether". Science. 297 (5585): 1330–3. Bibcode:2002Sci...297.1330C. doi:10.1126/science.1074611. PMID 12154198.
- ↑ Ornitz DM, Itoh N (2001). "Fibroblast growth factors". Genome Biology. 2 (3): reviews3005.1–reviews3005.12. doi:10.1186/gb-2001-2-3-reviews3005. PMC 138918
 . PMID 11276432.
. PMID 11276432. - 1 2 Duchesne L, Tissot B, Rudd TR, Dell A, Fernig DG (September 2006). "N-glycosylation of fibroblast growth factor receptor 1 regulates ligand and heparan sulfate co-receptor binding". The Journal of Biological Chemistry. 281 (37): 27178–89. doi:10.1074/jbc.M601248200. PMID 16829530.
- ↑ Coutts JC, Gallagher JT (December 1995). "Receptors for fibroblast growth factors". Immunology and Cell Biology. 73 (6): 584–9. doi:10.1038/icb.1995.92. PMID 8713482.
- ↑ Sleeman M, Fraser J, McDonald M, Yuan S, White D, Grandison P, Kumble K, Watson JD, Murison JG (June 2001). "Identification of a new fibroblast growth factor receptor, FGFR5". Gene. 271 (2): 171–82. doi:10.1016/S0378-1119(01)00518-2. PMID 11418238.
- ↑ Robinson CJ, Stringer SE (March 2001). "The splice variants of vascular endothelial growth factor (VEGF) and their receptors". Journal of Cell Science. 114 (Pt 5): 853–65. PMID 11181169.
- ↑ Myers SM, Eng C, Ponder BA, Mulligan LM (November 1995). "Characterization of RET proto-oncogene 3' splicing variants and polyadenylation sites: a novel C-terminus for RET". Oncogene. 11 (10): 2039–45. PMID 7478523.
- ↑ Baloh RH, Enomoto H, Johnson EM, Milbrandt J (February 2000). "The GDNF family ligands and receptors - implications for neural development". Current Opinion in Neurobiology. 10 (1): 103–10. doi:10.1016/S0959-4388(99)00048-3. PMID 10679429.
- ↑ Airaksinen MS, Titievsky A, Saarma M (May 1999). "GDNF family neurotrophic factor signaling: four masters, one servant?". Molecular and Cellular Neurosciences. 13 (5): 313–25. doi:10.1006/mcne.1999.0754. PMID 10356294.
- ↑ Arighi E, Borrello MG, Sariola H (2005). "RET tyrosine kinase signaling in development and cancer". Cytokine & Growth Factor Reviews. 16 (4-5): 441–67. doi:10.1016/j.cytogfr.2005.05.010. PMID 15982921.
- ↑ Fu HL, Valiathan RR, Arkwright R, Sohail A, Mihai C, Kumarasiri M, Mahasenan KV, Mobashery S, Huang P, Agarwal G, Fridman R (March 2013). "Discoidin domain receptors: unique receptor tyrosine kinases in collagen-mediated signaling". The Journal of Biological Chemistry. 288 (11): 7430–7. doi:10.1074/jbc.R112.444158. PMC 3597784
 . PMID 23335507.
. PMID 23335507.
External links
- Tyrosine Kinase Receptors at the US National Library of Medicine Medical Subject Headings (MeSH)
- EC 2.7.10.1