Active site
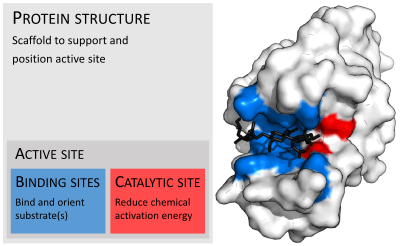
In biology, the active site is the region of an enzyme where substrate molecules bind and undergo a chemical reaction. The active site consists of residues that form temporary bonds with the substrate (binding site) and residues that catalyse a reaction of that substrate (catalytic site). The active site is usually a groove or pocket of the enzyme which can be located in a deep tunnel within the enzyme,[1] or between the interfaces of multimeric enzymes. An active site can catalyse a reaction repeatedly as its residues are not altered at the end of the reaction (they may change during the reaction, but are regenerated by the end).[2]
Binding site


Usually, an enzyme molecule has only one active site, and the active site fits with one specific type of substrate. An active site contains a binding site that binds the substrate and orients it for catalysis. Residues in the binding site form hydrogen bonds, hydrophobic interactions, or temporary covalent interactions (van der Waals) with the substrate to make an enzyme-substrate complex. In order to function, the active site needs to be in a specific conformation and so denaturation of the protein by high temperatures or extreme pH values will destroy its catalytic activity.[3] A tighter fit between an active site and the substrate molecule is believed to increase efficiency of a reaction.[4] Most enzymes have deeply buried active sites, which can be accessed by a substrate via access channels.[1]
There are two proposed models of how enzymes fit to their specific substrate: the lock and key model and the induced fit model.
Lock and key hypothesis
Emil Fischer's lock and key model assumes that the active site is a perfect fit for a specific substrate and that once the substrate binds to the enzyme no further modification occurs.[5]
Induced fit hypothesis
Daniel Koshland's theory of enzyme-substrate binding is that the active site and the binding portion of the substrate are not exactly complementary.[5] The induced fit model is a development of the lock-and-key model and assumes that an active site is flexible and it changes shape until the substrate is completely bound. The substrate is thought to induce a change in the shape of the active site. The hypothesis also predicts that the presence of certain residues (amino acids) in the active site will encourage the enzyme to locate the correct substrate. Conformational changes may then occur as the substrate is bound. After the products of the reaction move away from the enzyme, the active site returns to its initial shape.
Catalytic site
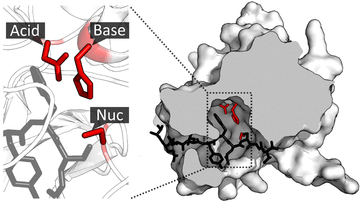
Once the substrate is bound and oriented in the active site, catalysis can begin. The residues of the catalytic site are typically very close to the binding site, and some residues can have dual-roles in both binding and catalysis.
Catalytic residues of the site interact with the substrate to lower the activation energy of a reaction and so make it proceed faster. They do this by a number of different mechanisms. Firstly, they can act as donors or acceptors of protons or other groups on the substrate to facilitate the reaction. They can also form electrostatic interactions to stabilise charge buildup on the transition state or leaving group. They can also directly take part in covalent catalysis, forming an acyl-enzyme intermediate that is then resolved to regenerate the catalytic residue and release the product. In this way, the catalytic residues of the active site provide an alternative reaction mechanism with lower activation energy.
Cofactors
Enzymes can use cofactors as ‘helper molecules’. Coenzymes are one example of cofactors. Coenzymes bind to the enzyme temporarily and are released after the reaction has occurred. Metal ions are another type of cofactor.
Inhibitors
Inhibitors disrupt the interaction between enzyme and substrate, slowing down the rate of a reaction. There are different types of inhibitor, including both reversible and irreversible forms. Reversible inhibitors can be competitive or non-competitive. Competitive reversible inhibitors have a similar shape to the substrate and bind to the enzyme’s active site temporarily, blocking entry of the actual substrate into the active site. Non-competitive reversible inhibitors bind to the enzyme however not in the active site. Despite not interacting with the active site, non-competitive inhibitors do reduce the rate of the reaction because they cause the enzyme to change shape. Irreversible inhibitors bind permanently to the enzyme, blocking access to active sites and therefore reducing the rate of the reaction.
| Example | Binds active site? | Reduces rate of reaction? | |
|---|---|---|---|
| Competitive reversible inhibitor | HIV protease inhibitors | Yes | Yes |
| Non-competitive reversible inhibitor | Heavy metals such as lead and mercury | No | Yes |
| Irreversible inhibitor | Cyanide | Yes | Yes |
Active sites in drug discovery
Identification of active sites is crucial in the process of drug discovery. The 3-D structure of the enzyme is analysed to identify active sites and design drugs which can fit into them. Proteolytic enzymes are targets for some drugs, such as protease inhibitors, which include drugs against AIDS and hypertension.[6] These protease inhibitors bind to an enzyme's active site and block interaction with natural substrates.[7] An important factor in drug design is the strength of binding between the active site and an enzyme inhibitor.[8]
Active sites can be mapped to aid design of new drugs such as enzyme inhibitors. This involves description of the size of an active site and the number and properties of sub-sites, such as details of the binding interaction.[6] Modern database technology called CPASS (Comparison of Protein Active Site Structures) however allows us to compare active sites in more detail and to look at structural similarity using software.[9]
Allosteric sites
An allosteric site is a site on an enzyme, unrelated to its active site, which can bind an effector molecule. This interaction is another mechanism of enzyme regulation. Allosteric modification usually happens in proteins with more than one subunit. Allosteric interactions are often present in metabolic pathways and are beneficial in that they allow one step of a reaction to regulate another step.[7] They allow an enzyme to have a range of molecular interactions, other than the highly specific active site.[7]
See also
Notes
References
- 1 2 Pravda L.; Berka K.; Svobodova Varekova R; Banas P.; Laskowski R.A.; Koca J.; Otyepka M. (2014). "Anatomy of Enzyme Channels". BMC Bioinformatics. 15: 379. doi:10.1186/s12859-014-0379-x.
- ↑ Alberts, B (2010). Essential Cell Biology. Garland Science. p. 91.
- ↑ Campbell, P (2006). Biochemistry Illustrated. Elsevier. pp. 83–85.
- ↑ Kool ET (1984). "Active site tightness and substrate fit in DNA replication". Annual Review of Biochemistry. 71: 191–219. doi:10.1146/annurev.biochem.71.110601.135453.
- 1 2 Sullivan SM (2008). "Enzymes with lid-gated active sites must operate by an induced fit mechanism instead of conformational selection". Proceedings of the National Academy of Sciences of the United States of America. 105: 13829–13834. doi:10.1073/pnas.0805364105.
- 1 2 Schechter I (2005). "Mapping of the active site of proteases in the 1960s and rational design of inhibitors/drugs in the 1990s". Current Protein and Peptide Science. 6: 501–512. doi:10.2174/138920305774933286.
- 1 2 3 DeDecker BS (2000). "Allosteric drugs: thinking outside the active-site box". Chemistry and Biology. 7: 103–107. doi:10.1016/S1074-5521(00)00115-0.
- ↑ Zuercher M (2008). "Structure-Based Drug Design: Exploring the Proper Filling of Apolar Pockets at Enzyme Active Sites". Journal of Organic Chemistry. 73: 4345–4361. doi:10.1021/jo800527n.
- ↑ Powers R (2006). "Comparison of protein active site structures for functional annotation of proteins and drug design". Proteins : Structure, Function, and Bioinformatics. 65: 124–135. doi:10.1002/prot.21092.
Further reading
- Alan Fersht, Structure and Mechanism in Protein Science : A Guide to Enzyme Catalysis and Protein Folding. W. H. Freeman, 1998. ISBN 0-7167-3268-8
- Bugg, T. Introduction to Enzyme and Coenzyme Chemistry. (2nd edition), Blackwell Publishing Limited, 2004. ISBN 1-4051-1452-5.
| Wikimedia Commons has media related to Active site. |
