Sphingomyelin phosphodiesterase
| Sphingomyelin phosphodiesterase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 Crystal structure of sphingomyelinase from Bacillus cereus.[1] | |||||||||
| Identifiers | |||||||||
| EC number | 3.1.4.12 | ||||||||
| CAS number | 9031-54-3 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / EGO | ||||||||
| |||||||||
Sphingomyelin phosphodiesterase (EC 3.1.4.12, also known as neutral sphingomyelinase, sphingomyelinase, or SMase) is a hydrolase enzyme that is involved in sphingolipid metabolism reactions. SMase is a member of the DNase I superfamily of enzymes and is responsible for breaking sphingomyelin (SM) down into phosphocholine and ceramide. The activation of SMase has been suggested as a major route for the production of ceramide in response to cellular stresses.[2]
Sphingomyelinase family
Currently, five types of SMase have been identified. These are classified according to their cation dependence and pH optima of action and are:
- Lysosomal acid SMase
- Secreted zinc-dependent acid SMase
- Magnesium-dependent neutral SMase
- Magnesium-independent neutral SMase
- Alkaline SMase
Of these, the lysosomal acidic SMase and the magnesium-dependent neutral SMase are considered major candidates for the production of ceramide in the cellular response to stress.
Neutral sphingomyelinase
Neutral sphingomyelinase (N-SMase) activity was first described in fibroblasts from patients with Niemann-Pick disease – a lysosomal storage disease characterized by deficiencies in acid SMase.[3] Subsequent study found that this enzyme was the product of a distinct gene, had an optimum pH of 7.4, was dependent on Mg2+ ions for activity, and was particularly enriched in brain.[4] However, a more recent study in bovine brain suggested the existence of multiple N-SMase isoforms with different biochemical and chromatographical properties.[5]
A major breakthrough came in the mid-1980s with the cloning of the first N-SMases from Bacillus cereus and Staphylococcus aureus.[6][7] Using the sequences of these bacterial sphingomyelinases in homology searches ultimately led to the identification of the yeast N-SMases ISC1 in the budding yeast Saccharomyces cerevisiae[8] and the mammalian N-SMase enzymes, nSMase1 and nSMase2.[9][10][11] The identity between mammalian, yeast and bacterial SMases is very low - being approximately 20% between nSMase2 and the B. cereus SMase. However, an alignment of the sequences (see figure) indicate a number of conserved residues throughout the family, particularly in the catalytic region of the enzymes.[12] This has led to the suggestion of a common catalytic mechanism for the N-SMase family.
A third N-SMase protein – termed nSMase3 – was recently cloned and characterized. nSMase3 bears little sequence similarity to either nSMase1 or nSMase2. However, there appears to be a high degree of evolutionary conservation from lower to higher organisms, suggesting that it may comprise a unique and distinct N-SMase. The high expression of nSMase3 in heart and skeletal muscle also suggests potential roles in heart function.[13]
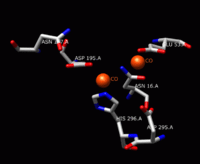
Active site
The solving of the crystal structure of the neutral sphingomyelinase from Listeria ivanovii and Bacillus cereus has allowed a fuller understanding of their enzymatic site. The active site of the B. cereus SMase comprises the residues Asn-16, Glu-53, Asp-195, Asn-197, and His-296. Of these, the residues Glu-53, Asp-195, and His-296 are known to be essential for activity. The relative catalytic activities of SMase when metal ions are bound to the active site have been studied for divalent metal ions Co2+, Mn2+, Mg2+, Ca2+, and Sr2+. Of these five metal ions, Co2+, Mn2+, and Mg2+ bound to the active site result in high catalytic activity of SMase. Ca2+ and Sr2+ bound to the active site exhibit much lower catalytic activity of SMase. When one Mg2+ ion or two Co2+ ions bind to the active site, double hexa-coordinated geometry results with two octahedral bi-pyramids for Co2+ and one octahedral bi-pyramid for Mg2+. When one Ca2+ ion binds to the active site, a hepta-coordinated geometry results. Therefore, the difference in catalytic activity for metal ions is predicted to be due to geometrical differences. Of Co2+ and Mg2+, SMase has better reactivity when two Co2+ ions are bound to SMase; when these Co2+ ions are bound, Glu-53 and His-296 each bind one divalent metal cation. These cations are surrounded by bridged water molecules and function as Lewis acids.[1]
Mechanism

The solving of the crystal structure of the neutral sphingomyelinase from Listeria ivanovii and Bacillus cereus has also shed light on their catalytic mechanisms. The active site of SMase contains Glu and His residues that are each bound to one or two divalent metal cations, usually Co2+, Mg2+, or Ca2+ for optimum performance. These two cations assist in catalysis by recruiting SM to the active site of SMase. The divalent cation bound to the Glu residue interacts with the amido-oxygen and ester-oxygen between C1 and the phosphate group of SM; an Asn residue and the divalent metal cation bound to the His residue bind to the oxygen atoms of the phosphate group of SM. This stabilizes the phosphate group’s negative charge. The metal cation bound to the His residue and Asp and Asn side chains lower the pKa value of one of the bridged water molecules, thus activating a water molecule. This water molecule then acts as a nucleophile and attacks the phosphate group of SM, creating a pentavalent phosphorus atom whose negative charge is stabilized by the divalent metal cations. The phosphate then reforms its tetrahedral conformation and results in the products ceramide and phosphocholine.[1] However, it is currently not clear if the mechanism of action of the acidic sphingomyelinase is the same, owing to the lack of a crystal structure.
References
- 1 2 3 PDB: 2ddt; Ago H, Oda M, Takahashi M, Tsuge H, Ochi S, Katunuma N, Miyano M, Sakurai J (June 2006). "Structural basis of the sphingomyelin phosphodiesterase activity in neutral sphingomyelinase from Bacillus cereus". J. Biol. Chem. 281 (23): 16157–67. doi:10.1074/jbc.M601089200. PMID 16595670.
- ↑ Hannun YA, Obeid LM (July 2002). "The Ceramide-centric universe of lipid-mediated cell regulation: stress encounters of the lipid kind". J. Biol. Chem. 277 (29): 25847–50. doi:10.1074/jbc.R200008200. PMID 12011103.
- ↑ Schneider PB, Kennedy EP (May 1967). "Sphingomyelinase in normal human spleens and in spleens from subjects with Niemann-Pick disease". J. Lipid Res. 8 (3): 202–9. PMID 4962590.
- ↑ Rao BG, Spence MW (September 1976). "Sphingomyelinase activity at pH 7.4 in human brain and a comparison to activity at pH 5.0". J. Lipid Res. 17 (5): 506–15. PMID 9463.
- ↑ Jung SY, Suh JH, Park HJ, Jung KM, Kim MY, Na DS, Kim DK (September 2000). "Identification of multiple forms of membrane-associated neutral sphingomyelinase in bovine brain". J. Neurochem. 75 (3): 1004–14. doi:10.1046/j.1471-4159.2000.0751004.x. PMID 10936181.
- ↑ Coleman DC, Arbuthnott JP, Pomeroy HM, Birkbeck TH (December 1986). "Cloning and expression in Escherichia coli and Staphylococcus aureus of the beta-lysin determinant from Staphylococcus aureus: evidence that bacteriophage conversion of beta-lysin activity is caused by insertional inactivation of the beta-lysin determinant". Microb. Pathog. 1 (6): 549–64. doi:10.1016/0882-4010(86)90040-9. PMID 3334158.
- ↑ Yamada A, Tsukagoshi N, Udaka S, Sasaki T, Makino S, Nakamura S, Little C, Tomita M, Ikezawa H (August 1988). "Nucleotide sequence and expression in Escherichia coli of the gene coding for sphingomyelinase of Bacillus cereus". Eur. J. Biochem. 175 (2): 213–20. doi:10.1111/j.1432-1033.1988.tb14186.x. PMID 2841128.
- ↑ Sawai H, Okamoto Y, Luberto C, Mao C, Bielawska A, Domae N, Hannun YA (December 2000). "Identification of ISC1 (YER019w) as inositol phosphosphingolipid phospholipase C in Saccharomyces cerevisiae". J. Biol. Chem. 275 (50): 39793–8. doi:10.1074/jbc.M007721200. PMID 11006294.
- ↑ Tripathi,K. "Role of Inositol Phosphosphingolipid Phospholipase C1, the Yeast Homolog of Neutral Sphingomyelinases in DNA Damage Response and Diseases.". J Lipids. 2015;2015:161392. doi:10.1155/2015/161392.
- ↑ Tomiuk S, Hofmann K, Nix M, Zumbansen M, Stoffel W (March 1998). "Cloned mammalian neutral sphingomyelinase: functions in sphingolipid signaling?". Proc. Natl. Acad. Sci. U.S.A. 95 (7): 3638–43. doi:10.1073/pnas.95.7.3638. PMC 19888
 . PMID 9520418.
. PMID 9520418. - ↑ Tomiuk S, Zumbansen M, Stoffel W (February 2000). "Characterization and subcellular localization of murine and human magnesium-dependent neutral sphingomyelinase". J. Biol. Chem. 275 (8): 5710–7. doi:10.1074/jbc.275.8.5710. PMID 10681556.
- ↑ Clarke CJ, Snook CF, Tani M, Matmati N, Marchesini N, Hannun YA (September 2006). "The extended family of neutral sphingomyelinases". Biochemistry. 45 (38): 11247–56. doi:10.1021/bi061307z. PMID 16981685.
- ↑ Krut O, Wiegmann K, Kashkar H, Yazdanpanah B, Krönke M (May 2006). "Novel tumor necrosis factor-responsive mammalian neutral sphingomyelinase-3 is a C-tail-anchored protein". J. Biol. Chem. 281 (19): 13784–93. doi:10.1074/jbc.M511306200. PMID 16517606.
Further reading
External links
- Sphingomyelin Phosphodiesterase at the US National Library of Medicine Medical Subject Headings (MeSH)