Viroplasm
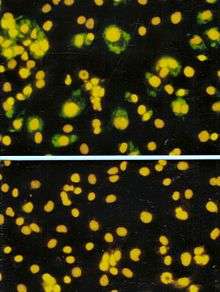
A viroplasm is an inclusion body in a cell where viral replication and assembly occurs. They may be thought of as viral factories in the cell. There are many viroplasms in one infected cell, where they appear dense to electron microscopy. Very little is understood about the mechanism of viroplasm formation.
Definition
A viroplasm is a perinuclear or a cytoplasmic large compartments where viral replication and assembly occurs.[1] The viroplasm formation is caused by the interactions between the virus and the infected cell, where viral products and cell elements are confined.[1] Virosomes are also called 'virus factories', and 'virus inclusions'.[2]
Groups of viruses that form viroplasms
Viroplasms have been reported in many unrelated groups of Eukaryotic viruses that replicate in cytoplasm, however, viroplasms from plants' viruses have not been as studied as viroplasms from animals' viruses.[1] Viroplasms have been found in the cauliflower mosaic virus,[3] rotavirus,[4] vaccinia virus[5] and the rice dwarf virus.[6] These appear electron-dense under an electron microscope and are insoluble.[1]
Structure and formation
Viroplasms are localized in the perinuclear area or in the cytoplasm of infected cells and are formed early in the infection cycle.[1][15] The number and the size of viroplasms depend on the virus, the virus isolate, hosts species, and the stage of the infection.[16] For example, viroplasms of mimivirus have a similar size to the nucleus of its host, the amoeba Acanthamoeba polyphaga.[9]
A virus can induce changes in composition and organization of host cell cytoskeletal and membrane compartments, depending on the step of the viral replication cycle.[2] This process involves a number of complex interactions and signaling events between viral and host cell factors.
Viroplasms are formed early during the infection; in many cases, the cellular rearrangements caused during virus infection lead to the construction of sophisticated inclusions —viroplasms— in the cell where the factory will be assembled. The viroplasm is where components such as replicase enzymes, virus genetic material, and host proteins required for replication concentrate, and thereby increase the efficiency of replication.[2] At the same time, large amounts of ribosomes, protein-synthesis components, protein folding chaperones, and mitochondria are recruited. Some of the membrane components are used for viral replication while some others will be modified to produce viral envelopes, when the viruses are enveloped. The viral replication, protein synthesis and assembly require a considerable amount of energy, provided by large clusters of mitochondria at the periphery of viroplasms. The virus factory is often enclosed by a membrane derived from the rough endoplasmic reticulum or by cytoskeletal elements.[1][15]
In animal cells, virus particles are gathered by the microtubule-dependent aggregation of toxic or misfolded protein near the microtubule organizing center (MTOC), so the viroplasms of animal viruses are generally localized near the MTOC.[1][17] MTOCs are not found in plant cells. Plant viruses induce the rearrangement of membranes structures to form the viroplasm. This is mostly shown for plant RNA viruses.[15]
Functions
Viroplasm is the location within the infected cell where viral replication and assembly take place.[1] Wrapping the viroplasm with a membrane, concentrates the viral components required for the genome replication and the morphogenesis of new virus particles, so it increases the efficiency of the processes.[1] The recruitment of cellular membranes and cytoskeleton to generate virus replication sites can also benefit viruses in other ways. Disruption of cellular membranes can, for example, slow the transport of immunomodulatory proteins to the surface of infected cells and protect against innate and acquired immune responses, and rearrangements to cytoskeleton can facilitate virus release.[2] The viroplasm could also prevent virus degradation by proteases and nucleases.[15]
In the case of the Cauliflower mosaic virus (CaMV), viroplasms improve the virus transmission by the aphid vector. Viroplasms also control release of virons when the insect stings an infected plant cell or a cell near the infected cells.[14]
Possible co-evolution with the host
Aggregated structures may protect viral functional complexes from the cellular degradation systems. For example, formation of viral factories of the ASFV viroplasm is very similar to the aggresome formation.[1] An aggresome is a perinuclear site where misfolded proteins are transported and stored by the cell components for their destruction. It has been proposed that the viroplasm could be the product of a co-evolution between the virus and its host.[14] It is possible that a cellular response originally designed to reduce the toxicity of misfolded proteins is exploited by cytoplasmic viruses to improve their replication, the virus capsid synthesis, and assembly.[14] Alternatively, the activation of host defense mechanisms may involve sequestration of virus components in aggregates to prevent their dissemination, followed by their neutralisation. For example, viroplasms of mammalian viruses contain certain elements of the cellular degradation machinery which might enable cellular protective mechanisms against viral components.[18] Given the co-evolution of viruses with their host cells, changes in cell structure induced during infection are likely to involve a combination of the two strategies.[1]
Use in diagnostics
Presence of viroplasms is used to diagnose certain viral infections. Understanding the phenomena of virus aggregation and of the cell response to the presence of virus, and whether viroplasms facilitate or inhibit viral replication, may help to develop new therapeutic approaches against virus infections in animal and plant cells.[15]
See also
References
- 1 2 3 4 5 6 7 8 9 10 11 12 13 14 Novoa, R. R.; Calderita, G.; Arranz, R.; Fontana, J.; Granzow, H.; Risco, C. (Feb 2005). "Virus factories: associations of cell organelles for viral replication and morphogenesis". Biol Cell. 97 (2): 147–72. doi:10.1042/bc20040058.
- 1 2 3 4 Netherton, Christopher; Moffat, Kathy; Brooks, Elizabeth; Wileman, Thomas (2007). "A Guide to Viral Inclusions, Membrane Rearrangements, Factories, and Viroplasm Produced During Virus Replication". Advances in Virus Research. 70: 101–182. doi:10.1016/S0065-3527(07)70004-0. Retrieved 2015-09-23.
- ↑ Xiong; Muller, S; Lebeurier, G; Hirth, L (1982). "Identification by immunoprecipitation of cauliflower mosaic virus in vitro major translation product with a specific serum against viroplasm protein". The EMBO Journal. 1 (8): 971–976. PMC 553144
 . PMID 16453427.
. PMID 16453427. - ↑ Nilsson; Von Bonsdorff, CH; Weclewicz, K; Cohen, J; Svensson, L (1998). "Assembly of viroplasm and virus-like particles of rotavirus by a Semliki Forest virus replicon". Virology. 242 (2): 255–65. doi:10.1006/viro.1997.8987. PMID 9514960.
- ↑ Szajner; Weisberg, AS; Wolffe, EJ; Moss, B (2001). "Vaccinia virus A30L protein is required for association of viral membranes with dense viroplasm to form immature virions". Journal of Virology. 75 (13): 5752–61. doi:10.1128/JVI.75.13.5752-5761.2001. PMC 114291
 . PMID 11390577.
. PMID 11390577. - ↑ Wei; Kikuchi, A; Suzuki, N; Shimizu, T; Hagiwara, K; Chen, H; Omura, T (2006). "Pns4 of rice dwarf virus is a phosphoprotein, is localized around the viroplasm matrix, and forms minitubules". Archives of Virology. 151 (9): 1701–12. doi:10.1007/s00705-006-0757-4. PMID 16609816.
- ↑ Sodeik, Beate; Doms, Robert W.; Ericsson, Maria; Hiller, Gerhard; Machamer, Carolyn E.; Wouter; Moss, Bernard; Grittiths, Gareth (1993). "Assembly of Vaccinia Virus: Role of the Intermediate Compartment Between the Endoplasmic Reticulum and the Golgi Stacks". The Journal of Cell Biology. 121 (3): 521–541. doi:10.1083/jcb.121.3.521.
- ↑ Salas, María L.; Andrés, Germán (2013). "African swine fever virus morphogenesis". Virus Research. 173: 29–41. doi:10.1016/j.virusres.2012.09.016.
- 1 2 Suzan-Monti, Marie; La Scola, Bernard; Barrassi, Lina; Espinosa, Leon; Raoult, Didier. "Ultrastructural Characterization of the Giant Volcano-like Virus Factory of Acanthamoeba polyphaga Mimivirus". PLoS ONE. 2 (3): e328. doi:10.1371/journal.pone.0000328.

- ↑ Fernando; Costas, Javier Benavente (2004). "374Avian Reovirus Morphogenesis Occurs Within Viral Factories and Begins with the Selective Recruitment of σNS and λA to μNS Inclusions .". Journal of Molecular Biology. 341 (2): 361–374. doi:10.1016/j.jmb.2004.06.026.
- ↑ Fontana, J; López-Iglesias, C; Tzeng, WP; Frey, TK; Fernández, JJ; Risco, C (Sep 2010). "Three-dimensional structure of Rubella virus factories". Virology. 405 (2): 579–91. doi:10.1016/j.virol.2010.06.043.
- ↑ Harper, Francis; Gaudin, Yves; Danielle; Xavier Lahaye, Blondel; Vidy, Aurore; Pomier, Carole; Obiang, Linda (2009). "Transcription and Replication Evidence that NBs Are Sites of Viral (NBs) in Rabies Virus-Infected Cells: Functional Characterization of Negri Bodies". J. Virol. 83 (16): 7948–7958. doi:10.1128/JVI.00554-09.
- ↑ Karczewski, M K; Strebel, K (1996). "Cytoskeleton association and virion incorporation of the human immunodeficiency virus type 1 Vif protein". J. Virol. 70 (1): 494–507.
- 1 2 3 4 Bak A., Gargani D., Macia J-L., Malouvet E., Vernerey M_S., Blanc S. and Drucker,M. Virus factories of Cauliflower mosaic virus are virion reservoirs that engage actively in vector-transmission. 2013 journal of Virology
- 1 2 3 4 5 Moshe, A.; Gorovits, R. (2012). "Virus-Induced Aggregates in Infected Cells". Viruses. 4: 2218–2232. doi:10.3390/v4102218.
- ↑ Shalla, TA; Shepherd, RJ; Peterson, LJ. "Comparative cytology of nine isolates of Cauliflower mosaic virus". 1980". Virology. 102: 381–388. doi:10.1016/0042-6822(80)90105-1.
- ↑ Wileman, T (2006). "Aggresomes and autophagy generate sites for viral infection". Science. 312: 875–878. doi:10.1126/science.1126766. PMID 16690857.
- ↑ Kopito, R (2000). "Aggresomes, inclusion bodies and protein aggregation". Trends Cell Biol. 10: 524–530. doi:10.1016/s0962-8924(00)01852-3.