Variant surface glycoprotein
| Variable surface glycoprotein | |
|---|---|
| Identifiers | |
| Organism | |
| Symbol | Tb927.5.4730 |
| Alt. symbols | Tb05.26C7.380 |
| Entrez | 3657576 |
| Other data | |
| Chromosome | 5: 1.41 - 1.41 Mb |
| Variant surface glycoprotein MITAT 1.2 | |
|---|---|
| Identifiers | |
| Organism | |
| Symbol | N/A |
| Alt. symbols | VSG 221 |
| UniProt | P26332 |
Variant surface glycoproteins (VSG) are a type of proteins present on the cell surface of parasitic protozoans belonging to the genus Trypanosoma. They are discovered from Trypanosoma brucei. They are used by the parasites to evade the host's immune system by mean of antigenic variation.
Combinatorial processes increase the diversity of variable surface glycoproteins.[1] The parasite is expressing a series of antigenically distinct VSGs from more than 2000 complete and partial (pseudogenes) VSG genes. The genes are located in telomeric and subtelomeric regions and are often activated by the duplicative transposition of a silent basic copy gene into an unlinked telomerically located expression site, producing an active expression-linked copy of that gene.[2]
In Trypanosoma brucei
In Trypanosoma brucei, the cell surface is covered by a dense coat of ~5 x 106 molecules of variant surface glycoprotein.[3] These glycoproteins act as major surface antigens.[4]
The two properties of the VSG coat that allow immune evasion are:
- Shielding — the dense nature of the VSG coat prevents the immune system of the mammalian host from accessing the plasma membrane or any other invariant surface epitopes (such as ion channels, transporters, receptors etc.) of the parasite. The coat is uniform, made up of millions of copies of the same molecule; therefore the only parts of the trypanosome the immune system can 'see' are the N-terminal loops of the VSG that make up the coat.[5]
- Periodic antigenic variation — the VSG coat undergoes frequent stochastic genetic modification — 'switching' — allowing variants expressing a new VSG coat to escape the specific immune response raised against the previous coat.
VSGs are replaced by an equally dense coat of procyclins when the parasite differentiates into the procylic form in the tsetse fly midgut. There is a very fast inhibition of VSG gene transcription which occurs as soon as the temperature is lowered.[6]
The VSGs from T. brucei are attached to the plasma membrane via a glycosyl-phosphatidylinositol (GPI) anchor.[7]
The coat is composed of VSG dimers and forms a macromolecular diffusion barrier. VSGs vary in primary amino acid sequence, the number of N-glycosylation sites, and the types of N-linked oligosaccharides and glycosylphosphatidylinositol membrane anchors they contain. VSG MITat.1.5 is glycosylated at all three potential N-glycosylation sites.[8]
Structure
VSG genes are hugely variable at the sequence level, but variants are thought to have strongly conserved structural features, based on two determined 3-dimensional structures and conservation of 2-dimensional sequence motifs, allowing them to perform a similar shielding function.[9] VSGs are made up of a highly variable N terminal domain of around 300 to 350 amino acids, and a more conserved C terminal domain of around 100 amino acids. N-terminal domains dimerise to form a bundle of four alpha helices, around which hang smaller structural features. VSG is anchored to the cell membrane via a glycophosphatidylinositol (GPI) anchor—a covalent linkage from the C-terminus, to approximately four sugars, to a phosphatidylinositol phospholipid acid which lies in the cell membrane.
The source of VSG variability during infection is a large 'archive' of VSG genes present in the T. brucei genome. Some of these are full-length, intact genes; others are pseudogenes) typically with frameshift mutations, premature stop codons, or fragmentation.[10] Expression of an antigenically different VSG can occur by simply switching to a different full-length VSG gene. In addition, chimeric or 'mosaic' VSG genes can be generated by combining segments from more than one silent VSG gene. The formation of mosaic VSGs allows the (partial) expression of pseudogene VSGs, which can constitute the major portion of the VSG archive, and can contribute directly to antigenic variation, vastly increasing the trypanosome's capacity for immune evasion and potentially posing a major problem for vaccine development.[11]
Antigenic variation
VSG is highly immunogenic, and an immune response raised against a specific VSG coat will rapidly kill trypanosomes expressing this variant. Antibody-mediated trypanosome killing can also be observed in vitro by a complement-mediated lysis assay. However, with each cell division there is a possibility that one or both of the progeny will switch expression to change the VSG that is being expressed. The frequency of VSG switching has been measured to be approximately 0.1% per division.[12] As T. brucei populations can peak at a size of 1011 within a host [13] this rapid rate of switching ensures that the parasite population is constantly diverse. A diverse range of coats expressed by the trypanosome population means that the immune system is always one step behind: it takes several days for an immune response against a given VSG to develop, giving the population time to diversify as individuals undergo further switching events. Reiteration of this process prevents extinction of the infecting trypanosome population, allowing chronic persistence of parasites in the host, enhancing opportunities for transmission. The clinical effect of this cycle is successive 'waves' of parasitaemia (trypanosomes in the blood).[3]
Expression
The source of VSG variability during infection is a large 'archive' of VSG genes present in the T. brucei genome. Some of these are full-length, intact genes; others are pseudogenes) typically with frameshift mutations, premature stop codons, or fragmentation.[10] Expression of an antigenically different VSG can occur by simply switching to a different full-length VSG gene. In addition, chimeric or 'mosaic' VSG genes can be generated by combining segments from more than one silent VSG gene. The formation of mosaic VSGs allows the (partial) expression of pseudogene VSGs, which can constitute the major portion of the VSG archive, and can contribute directly to antigenic variation, vastly increasing the trypanosome's capacity for immune evasion and potentially posing a major problem for vaccine development.[11]
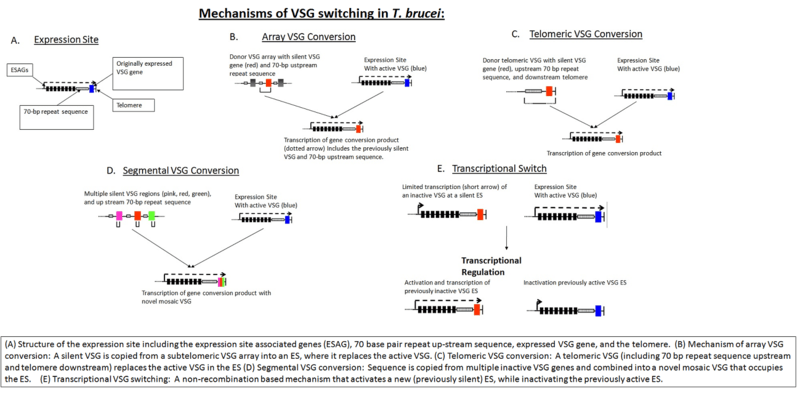
VSG genes can be kept silent and switched on at a given time. The expressed VSG is always located in an Expression Site (ES), which are specialised expression loci found at the telomeres of some of the large and intermediate chromosomes. Each ES is a polycistronic unit, containing a number of Expression Site-Associated Genes (ESAGs) all expressed along with the active VSG. While multiple ES exist, only a single one is ever active at one time. A number of mechanisms appear to be involved in this process, but the exact nature of the silencing is still unclear.[14]
The expressed VSG can be switched either by activating a different expression site (and thus changing to express the VSG in that site), or by changing the VSG gene in the active site to a different variant. The genome contains many copies of VSG genes, both on minichromosomes and in repeated sections in the interior of the chromosomes. These are generally silent, typically with omitted sections or premature stop codons, but are important in the evolution of new VSG genes. It is estimated up to 10% of the T.brucei genome may be made up of VSG genes or pseudogenes. Any of these genes can be moved into the active site by recombination for expression. Again, the exact mechanisms that control this are unclear, but the process seems to rely on DNA repair machinery and a process of homologous recombination.[15]
The bloodstream expression site (BES), or telomeric expression site, is used for exchanging variant surface glycoproteins when in host's blood stream to escape the complement system. BESs are polymorphic in size and structure but reveal a surprisingly conserved architecture in the context of extensive recombination. Very small BESs do exist and many functioning BESs do not contain the full complement of expression site associated genes (ESAGs).[16] There is a collection of an estimated 20-30 sites, each being active at a time.[17] Active VSG expression sites are depleted of nucleosomes.[18]
The gene repertoires in T. brucei have diverged to become strain-specific.[19]
The variant surface glycoprotein genes of T. brucei have been classified into two groups depending upon whether or not duplication of the genes is observed when they are expressed.[20]
In other trypanosomes
Variable surface glycoproteins are also found in other Trypanosoma species,
In Trypanosoma equiperdum, a parasite causing the covering sickness in horses, These proteins allow the parasite to efficiently evade the host animal's immune system.[21] These VSGs allow the organism to constantly manipulate and change the surface structure of its proteins, which means it is constantly being presented to the immune system as a new foreign organism and this prevents the body from mounting a large enough immune response to eradicate the disease.[21] In this sense, Trypanosoma equiperdum is a very efficient organism; it may infect less species than other diseases, but it infects and survives very efficiently within its specified hosts. The VSG proteins in T. equiperdum are also phosphorylated.[22]
A VSG gene from Trypanosoma evansi, a parasite that causes a form of surra in animals, has been cloned in Escherichia coli. The expressed protein is immunoreactive with all the sera combinations. The animals immunized with whole cell lysate or recombinant protein show similar antibody reactions in ELISA (Enzyme-linked immunosorbent assay) and CATT (card agglutination test for Trypanosomiasis).[23] The variable surface glycoprotein RoTat 1.2 PCR can be used as a specific diagnostic tool for the detection of T. evansi infections.[24]
The smallest VSG protein (40 kDa in size) to date (1996) has been found in Trypanosoma vivax, which bears little carbohydrate.[25]
In Trypanosoma congolense, in vitro analyses of the incorporated sugars after hydrolysis of the glycoprotein showed that glucosamine and mannose are utilized in the biosynthesis of the carbohydrate moiety directly whereas galactose was converted possibly to other intermediates before being incorporated into the antigen. The unglycosylated VSG with a molecular weight of 47 kDa had completely lost its size heterogeneity.[26]
See also
- Coat protein (disambiguation)
- Glycocalyx
- List of MeSH codes (D23)
- List of MeSH codes (D12.776.395)
- List of MeSH codes (D12.776.543)
- Amastin, another surface (trans-membrane) glycoprotein in trypanosomatid parasites[27]
References
- ↑ Thon G, Baltz T, Giroud C, Eisen H (Aug 1990). "Trypanosome variable surface glycoproteins: composite genes and order of expression". Genes & Development. 4 (8): 1374–83. doi:10.1101/gad.4.8.1374. PMID 2227415.
- ↑ Buck GA, Jacquemot C, Baltz T, Eisen H (Dec 1984). "Re-expression of an inactivated variable surface glycoprotein gene in Trypanosoma equiperdum". Gene. 32 (3): 329–36. doi:10.1016/0378-1119(84)90008-8. PMID 6530143.
- 1 2 Barry JD, McCulloch R (2001). "Antigenic variation in trypanosomes: enhanced phenotypic variation in a eukaryotic parasite". Advances in Parasitology. 49: 1–70. doi:10.1016/S0065-308X(01)49037-3. ISBN 978-0-12-031749-3. PMID 11461029.
- ↑ Ferguson MA, Homans SW (Sep 1988). "Parasite glycoconjugates: towards the exploitation of their structure". Parasite Immunology. 10 (5): 465–79. doi:10.1111/j.1365-3024.1988.tb00236.x. PMID 3057422.
- ↑ Overath P, Chaudhri M, Steverding D, Ziegelbauer K (Feb 1994). "Invariant surface proteins in bloodstream forms of Trypanosoma brucei". Parasitology Today. 10 (2): 53–8. doi:10.1016/0169-4758(94)90393-X. PMID 15275499.
- ↑ Pays E, Coquelet H, Pays A, Tebabi P, Steinert M (Sep 1989). "Trypanosoma brucei: posttranscriptional control of the variable surface glycoprotein gene expression site". Molecular and Cellular Biology. 9 (9): 4018–21. PMC 362464
 . PMID 2779574.
. PMID 2779574. - ↑ Grab DJ, Verjee Y. "Localization of a Variable Surface Glycoprotein Phosphatidylinositol-Specific Phospholipase-C in Trypanosoma brucei brucei". FAO Corporate document depository. Food and Agricultural Organization of the United Nations.
- ↑ Mehlert A, Bond CS, Ferguson MA (Oct 2002). "The glycoforms of a Trypanosoma brucei variant surface glycoprotein and molecular modeling of a glycosylated surface coat". Glycobiology. 12 (10): 607–12. doi:10.1093/glycob/cwf079. PMID 12244073.
- ↑ Blum ML, Down JA, Gurnett AM, Carrington M, Turner MJ, Wiley DC (Apr 1993). "A structural motif in the variant surface glycoproteins of Trypanosoma brucei". Nature. 362 (6421): 603–9. doi:10.1038/362603a0. PMID 8464512.
- 1 2 Marcello L, Barry JD (Sep 2007). "Analysis of the VSG gene silent archive in Trypanosoma brucei reveals that mosaic gene expression is prominent in antigenic variation and is favored by archive substructure". Genome Research. 17 (9): 1344–52. doi:10.1101/gr.6421207. PMC 1950903
 . PMID 17652423.
. PMID 17652423. - 1 2 Barbour AG, Restrepo BI (2000). "Antigenic variation in vector-borne pathogens". Emerging Infectious Diseases. 6 (5): 449–57. doi:10.3201/eid0605.000502. PMC 2627965
 . PMID 10998374.
. PMID 10998374. - ↑ Turner CM (Aug 1997). "The rate of antigenic variation in fly-transmitted and syringe-passaged infections of Trypanosoma brucei". FEMS Microbiology Letters. 153 (1): 227–31. doi:10.1111/j.1574-6968.1997.tb10486.x. PMID 9252591.
- ↑ Barry JD, Hall JP, Plenderleith L (Sep 2012). "Genome hyperevolution and the success of a parasite". Annals of the New York Academy of Sciences. 1267 (1): 11–7. doi:10.1111/j.1749-6632.2012.06654.x. PMC 3467770
 . PMID 22954210.
. PMID 22954210. - ↑ Pays E (Nov 2005). "Regulation of antigen gene expression in Trypanosoma brucei". Trends in Parasitology. 21 (11): 517–20. doi:10.1016/j.pt.2005.08.016. PMID 16126458.
- ↑ Morrison LJ, Marcello L, McCulloch R (Dec 2009). "Antigenic variation in the African trypanosome: molecular mechanisms and phenotypic complexity". Cellular Microbiology. 11 (12): 1724–34. doi:10.1111/j.1462-5822.2009.01383.x. PMID 19751359.
- ↑ Hertz-Fowler C, Figueiredo LM, Quail MA, Becker M, Jackson A, Bason N, Brooks K, Churcher C, Fahkro S, Goodhead I, Heath P, Kartvelishvili M, Mungall K, Harris D, Hauser H, Sanders M, Saunders D, Seeger K, Sharp S, Taylor JE, Walker D, White B, Young R, Cross GA, Rudenko G, Barry JD, Louis EJ, Berriman M (2008). "Telomeric expression sites are highly conserved in Trypanosoma brucei". PloS One. 3 (10): e3527. Bibcode:2008PLoSO...3.3527H. doi:10.1371/journal.pone.0003527. PMC 2567434
 . PMID 18953401.
. PMID 18953401. - ↑ Vanhamme L, Lecordier L, Pays E (May 2001). "Control and function of the bloodstream variant surface glycoprotein expression sites in Trypanosoma brucei". International Journal for Parasitology. 31 (5-6): 523–31. doi:10.1016/S0020-7519(01)00143-6. PMID 11334937.
- ↑ Stanne TM, Rudenko G (Jan 2010). "Active VSG expression sites in Trypanosoma brucei are depleted of nucleosomes". Eukaryotic Cell. 9 (1): 136–47. doi:10.1128/EC.00281-09. PMC 2805301
 . PMID 19915073.
. PMID 19915073. - ↑ Hutchinson OC, Picozzi K, Jones NG, Mott H, Sharma R, Welburn SC, Carrington M (2007). "Variant Surface Glycoprotein gene repertoires in Trypanosoma brucei have diverged to become strain-specific". BMC Genomics. 8: 234. doi:10.1186/1471-2164-8-234. PMC 1934917
 . PMID 17629915.
. PMID 17629915. - ↑ Young JR, Turner MJ, Williams RO (1984). "The role of duplication in the expression of a variable surface glycoprotein gene of Trypanosoma brucei". Journal of Cellular Biochemistry. 24 (3): 287–95. doi:10.1002/jcb.240240309. PMID 6736139.
- 1 2 Raibaud A, Gaillard C, Longacre S, Hibner U, Buck G, Bernardi G, Eisen H (Jul 1983). "Genomic environment of variant surface antigen genes of Trypanosoma equiperdum". Proceedings of the National Academy of Sciences of the United States of America. 80 (14): 4306–10. Bibcode:1983PNAS...80.4306R. doi:10.1073/pnas.80.14.4306. PMC 384026
 . PMID 6308614.
. PMID 6308614. - ↑ Baltz T, Giroud C, Baltz D, Duvillier G, Degand P, Demaille J, Pautrizel R (1982). "The variable surface glycoproteins of Trypanosoma equiperdum are phosphorylated". The EMBO Journal. 1 (11): 1393–8. PMC 553222
 . PMID 6821334.
. PMID 6821334. - ↑ Sengupta PP, Balumahendiran M, Balamurugan V, Rudramurthy GR, Prabhudas K (Jun 2012). "Expressed truncated N-terminal variable surface glycoprotein (VSG) of Trypanosoma evansi in E. coli exhibits immuno-reactivity". Veterinary Parasitology. 187 (1-2): 1–8. doi:10.1016/j.vetpar.2012.01.012. PMID 22277627.
- ↑ Claes F, Radwanska M, Urakawa T, Majiwa PA, Goddeeris B, Büscher P (Sep 2004). "Variable Surface Glycoprotein RoTat 1.2 PCR as a specific diagnostic tool for the detection of Trypanosoma evansi infections". Kinetoplastid Biology and Disease. 3 (1): 3. doi:10.1186/1475-9292-3-3. PMC 521498
 . PMID 15377385.
. PMID 15377385. - ↑ Gardiner PR, Nene V, Barry MM, Thatthi R, Burleigh B, Clarke MW (Nov 1996). "Characterization of a small variable surface glycoprotein from Trypanosoma vivax". Molecular and Biochemical Parasitology. 82 (1): 1–11. doi:10.1016/0166-6851(96)02687-4. PMID 8943146.
- ↑ Reinwald E, Heidrich C, Risse HJ (May 1984). "In vitro studies on the biosynthesis of the surface glycoprotein of Trypanosoma congolense". The Journal of Protozoology. 31 (2): 300–6. doi:10.1111/j.1550-7408.1984.tb02966.x. PMID 6470988.
- ↑ Jackson AP (Jan 2010). "The evolution of amastin surface glycoproteins in trypanosomatid parasites". Molecular Biology and Evolution. 27 (1): 33–45. doi:10.1093/molbev/msp214. PMC 2794310
 . PMID 19748930.
. PMID 19748930.
External links
- Variant Surface Glycoproteins, Trypanosoma at the US National Library of Medicine Medical Subject Headings (MeSH)
- www.icp.ucl.ac.be