Fc receptor

An Fc receptor is a protein found on the surface of certain cells – including, among others, B lymphocytes, follicular dendritic cells, natural killer cells, macrophages, neutrophils, eosinophils, basophils, human platelets, and mast cells – that contribute to the protective functions of the immune system. Its name is derived from its binding specificity for a part of an antibody known as the Fc (Fragment, crystallizable) region. Fc receptors bind to antibodies that are attached to infected cells or invading pathogens. Their activity stimulates phagocytic or cytotoxic cells to destroy microbes, or infected cells by antibody-mediated phagocytosis or antibody-dependent cell-mediated cytotoxicity. Some viruses such as flaviviruses use Fc receptors to help them infect cells, by a mechanism known as antibody-dependent enhancement of infection.[1]
Classes of Fc receptor
There are several different types of Fc receptors (abbreviated FcR), which are classified based on the type of antibody that they recognize. The Latin letter used to identify a type of antibody is converted into the corresponding Greek letter, which is placed after the 'Fc' part of the name. For example, those that bind the most common class of antibody, IgG, are called Fc-gamma receptors (FcγR), those that bind IgA are called Fc-alpha receptors (FcαR) and those that bind IgE are called Fc-epsilon receptors (FcεR). The classes of FcR's are also distinguished by the cells that express them (macrophages, granulocytes, natural killer cells, T and B cells) and the signalling properties of each receptor.[2]
Fc-gamma receptors
All of the Fcγ receptors (FcγR) belong to the immunoglobulin superfamily and are the most important Fc receptors for inducing phagocytosis of opsonized (marked) microbes.[3] This family includes several members, FcγRI (CD64), FcγRIIA (CD32), FcγRIIB (CD32), FcγRIIIA (CD16a), FcγRIIIB (CD16b), which differ in their antibody affinities due to their different molecular structure.[4] For instance, FcγRI binds to IgG more strongly than FcγRII or FcγRIII does. FcγRI also has an extracellular portion composed of three immunoglobulin (Ig)-like domains, one more domain than FcγRII or FcγRIII has. This property allows FcγRI to bind a sole IgG molecule (or monomer), but all Fcγ receptors must bind multiple IgG molecules within an immune complex to be activated.[5]
The Fc-gamma receptors differ in their affinity for IgG and likewise the different IgG subclasses have unique affinities for each of the Fc gamma receptors.[6] These interactions are further tuned by the glycan (oligosaccharide) at position CH2-84.4 of IgG.[6] For example, by creating steric hindrance, fucose containing CH2-84.4 glycans reduce IgG affinity for FcγRIIIA.[6] In contrast, G0 glycans, which lack galactose and terminate instead with GlcNAc moieties, have increased affinity for FcγRIIIA.[6]
Another FcR is expressed on multiple cell types and is similar in structure to MHC class I. This receptor also binds IgG and is involved in preservation of this antibody.[7] However, since this Fc receptor is also involved in transferring IgG from a mother either via the placenta to her fetus or in milk to her suckling infant, it is called the neonatal Fc receptor (FcRn).[8][9] Recently, research suggested that this receptor plays a role in the homeostasis of IgG serum levels.
Fc-alpha receptors
Only one Fc receptor belongs to the FcαR subgroup, which is called FcαRI (or CD89).[10] FcαRI is found on the surface of neutrophils, eosinophils, monocytes, some macrophages (including Kupffer cells), and some dendritic cells.[10] It is composed of two extracellular Ig-like domains, and is a member of both the immunoglobulin superfamily and the multi-chain immune recognition receptor (MIRR) family.[3] It signals by associating with two FcRγ signaling chains.[10] Another receptor can also bind IgA, although it has higher affinity for another antibody called IgM.[11] This receptor is called the Fc-alpha/mu receptor (Fcα/μR) and is a type I transmembrane protein. With one Ig-like domain in its extracellular portion, this Fc receptor is also a member of the immunoglobulin superfamily.[12]
Fc-epsilon receptors
Two types of FcεR are known:[3]
- the high-affinity receptor FcεRI is a member of the immunoglobulin superfamily (it has two Ig-like domains). FcεRI is found on epidermal Langerhans cells, eosinophils, mast cells and basophils.[13][14] As a result of its cellular distribution, this receptor plays a major role in controlling allergic responses. FcεRI is also expressed on antigen-presenting cells, and controls the production of important immune mediators called cytokines that promote inflammation.[15]
- the low-affinity receptor FcεRII (CD23) is a C-type lectin. FcεRII has multiple functions as a membrane-bound or soluble receptor; it controls B cell growth and differentiation and blocks IgE-binding of eosinophils, monocytes, and basophils.[16]
Summary table
| Receptor name | Principal antibody ligand | Affinity for ligand | Cell distribution | Effect following binding to antibody |
| FcγRI (CD64) | IgG1 and IgG3 | High (Kd ~ 10−9 M) | Macrophages Neutrophils Eosinophils Dendritic cells |
Phagocytosis Cell activation Activation of respiratory burst Induction of microbe killing |
| FcγRIIA (CD32) | IgG | Low (Kd > 10−7 M) | Macrophages Neutrophils Eosinophils Platelets Langerhans cells |
Phagocytosis Degranulation (eosinophils) |
| FcγRIIB1 (CD32) | IgG | Low (Kd > 10−7 M) | B Cells Mast cells |
No phagocytosis Inhibition of cell activity |
| FcγRIIB2 (CD32) | IgG | Low (Kd > 10−7 M) | Macrophages Neutrophils Eosinophils |
Phagocytosis Inhibition of cell activity |
| FcγRIIIA (CD16a) | IgG | Low (Kd > 10−6 M) | NK cells Macrophages (certain tissues) |
Induction of antibody-dependent cell-mediated cytotoxicity (ADCC) Induction of cytokine release by macrophages |
| FcγRIIIB (CD16b) | IgG | Low (Kd > 10−6 M) | Eosinophils Macrophages Neutrophils Mast cells Follicular dendritic cells |
Induction of microbe killing |
| FcεRI | IgE | High (Kd ~ 10−10 M) | Mast cells Eosinophils Basophils Langerhans cells Monocytes |
Degranulation Phagocytosis |
| FcεRII (CD23) | IgE | Low (Kd > 10−7 M) | B cells Eosinophils Langerhans cells |
Possible adhesion molecule IgE transport across human intestinal epithelium Positive-feedback mechanism to enhance allergic sensitization (B cells) |
| FcαRI (CD89) | IgA | Low (Kd > 10−6 M) | Monocytes Macrophages Neutrophils Eosinophils |
Phagocytosis Induction of microbe killing |
| Fcα/μR | IgA and IgM | High for IgM, Mid for IgA | B cells Mesangial cells Macrophages |
Endocytosis Induction of microbe killing |
| FcRn | IgG | Monocytes Macrophages Dendritic cells Epithelial cells Endothelial cells Hepatocytes |
Transfers IgG from a mother to fetus through the placenta Transfers IgG from a mother to infant in milk Protects IgG from degradation |
Functions of Fc receptors
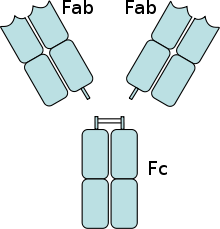
Fc receptors are found on a number of cells in the immune system including phagocytes like macrophages and monocytes, granulocytes like neutrophils and eosinophils, and lymphocytes of the innate immune system (natural killer cells) or adaptive immune system (e.g., B cells).[17][18][19] They allow these cells to bind to antibodies that are attached to the surface of microbes or microbe infected cells, helping these cells to identify and eliminate microbial pathogens. The Fc receptors bind the antibodies at their Fc region (or tail), an interaction that activates the cell that possesses the Fc receptor.[20] Activation of phagocytes is the most common function attributed to Fc receptors. For example, macrophages begin to ingest and kill an IgG-coated pathogen by phagocytosis following engagement of their Fcγ receptors.[21] Another process involving Fc receptors is called antibody-dependent cell-mediated cytotoxicity (ADCC). During ADCC, FcγRIII receptors on the surface of natural killer (NK) cells stimulate the NK cells to release cytotoxic molecules from their granules to kill antibody-covered target cells.[22] FcεRI has a different function. FcεRI is the Fc receptor on granulocytes, that is involved in allergic reactions and defense against parasitic infections. When an appropriate allergic antigen or parasite is present, the cross-linking of a least two of IgE molecules and their Fc receptors on the surface of a granulocyte will trigger the cell to rapidly release preformed mediators from its granules.[3]
Signaling mechanisms of Fc receptors
Fc gamma receptors generate signals within their cells through an important activation motif known as an Immunoreceptor tyrosine-based activation motif (ITAM).[23] An ITAM is a specific sequence of amino acids (YXXL) occurring twice in close succession in the intracellular tail of a receptor. When phosphate groups are added to the tyrosine (Y) residue of the ITAM by enzymes called tyrosine kinases, a signaling cascade is generated within the cell. This phosphorylation reaction typically follows interaction of an Fc receptor with its ligand. An ITAM is present in the intracellular tail of FcγRIIA, and its phosphorylation induces phagocytosis in macrophages. FcγRI and FcγRIIIA do not have an ITAM but can transmit an activating signal to their phagocytes by interacting with another protein that does. This adaptor protein is called the Fcγ subunit and, like FcγRIIA, contains the two YXXL sequences that are characteristic of an ITAM.
The presence of only one YXXL motif is not sufficient to activate cells, and represents a motif (I/VXXYXXL) known as an immunoreceptor tyrosine-based inhibitory motif (ITIM). FcγRIIB1 and FcγRIIB2 have an ITIM sequence and are inhibitory Fc receptors; they do not induce phagocytosis. Inhibitory actions of these receptors are controlled by enzymes that remove phosphate groups from tyrosine residues; the phosphatases SHP-1 and SHIP-1 inhibit signaling by Fcγ receptors.[24] Binding of ligand to FcγRIIB leads to phosphorylation of the tyrosine of the ITAM motif. This modification generates the binding site for the phosphatase, a SH2 recognition domain. The abrogation of ITAM activation signaling is caused by inhibition of protein tyrosine kinases of Src family, and by hydrolyzing the membrane PIP3 interrupting the further downstream signaling by the activating receptors, such as activating FcγRs, TCR, BCR and cytokine receptors (e.g. c-Kit).[25]
The negative signaling by FcγRIIB is mainly important for regulation of activated B cells. The positive B cell signaling is initiated by binding of foreign antigen to surface immunoglobulin. The same antigen-specific antibody is secreted and it can feedback-suppress, or promote negative signaling. This negative signaling is being provided by FcγRIIB.:[26] Experiments using B cell deletion mutants and dominant-negative enzymes have firmly established an important role for SH2-domain-containing inositol 5-phosphatase (SHIP) in negative signaling. Negative signaling through SHIP appears to inhibit the Ras pathway through SH2 domain competition with Grb2 and Shc and may involve consumption of intracellular lipid mediators that act as allosteric enzyme activators or that promote entry of extracellular Ca2+.[27]
Cellular activation by Fc Receptors
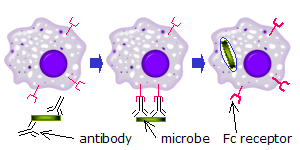
Fc receptors on phagocytes
When IgG molecules, specific for a certain antigen or surface component, bind to the pathogen with their Fab region (fragment antigen binding region), their Fc regions point outwards, in direct reach of phagocytes. Phagocytes bind those Fc regions with their Fc receptors.[21] Many low affinity interactions are formed between receptor and antibody that work together to tightly bind the antibody-coated microbe. The low individual affinity prevents Fc receptors from binding antibodies in the absence of antigen, and therefore reduces the chance of immune cell activation in the absence of infection. This also prevents agglutination (clotting) of phagocytes by antibody when there is no antigen. After a pathogen has been bound, interactions between the Fc region of the antibody and the Fc receptors of the phagocyte results in the initiation of phagocytosis. The pathogen becomes engulfed by the phagocyte by an active process involving the binding and releasing of the Fc region/Fc receptor complex, until the cell membrane of the phagocyte completely encloses the pathogen.[28]
Fc receptors on NK cells
The Fc receptor on NK cells recognize IgG that is bound to the surface of a pathogen-infected target cell and is called CD16 or FcγRIII.[29] Activation of FcγRIII by IgG causes the release of cytokines such as IFN-γ that signal to other immune cells, and cytotoxic mediators like perforin and granzyme that enter the target cell and promote cell death by triggering apoptosis. This process is known as antibody-dependent cell-mediated cytotoxicity (ADCC). FcγRIII on NK cells can also associate with monomeric IgG (i.e., IgG that is not antigen-bound). When this occurs, the Fc receptor inhibits the activity of the NK cell.[30]
Fc receptors on mast cells
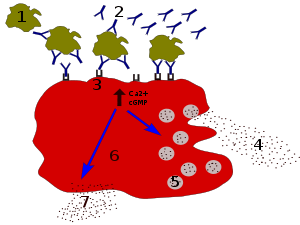
IgE antibodies bind to antigens of allergens. These allergen-bound IgE molecules interact with Fcε receptors on the surface of mast cells. Activation of mast cells following engagement of FcεRI results in a process called degranulation, whereby the mast cell releases preformed molecules from its cytoplasmic granules; these are a mixture of compounds including histamine, proteoglycans, and serine proteases.[31] Activated mast cells also synthesize and secrete lipid-derived mediators (such as prostaglandins, leukotrienes, and platelet-activating factor) and cytokines (such as interleukin 1, interleukin 3, interleukin 4, interleukin 5, interleukin 6, interleukin 13, tumor necrosis factor-alpha, GM-CSF, and several chemokines.[32][33] These mediators contribute to inflammation by attracting other leukocytes.
Fc receptors on eosinophils
Large parasites like the helminth (worm) Schistosoma mansoni are too large for ingestion by phagocytes. They also have an external structure called an integument that is resistant to attack by substances released by macrophages and mast cells. However, these parasites can become coated with IgE and recognized by FcεRI on the surface of eosinophils. Activated eosinophils release preformed mediators such as major basic protein, and enzymes such as peroxidase, against which helminths are not resistant.[34][35] The interaction of the FcεRI receptor with the Fc portion of helminth bound IgE causes the eosinophil to release these molecules in a mechanism similar to that of the NK cell during ADCC.[36]
See also
References
- ↑ Anderson R (2003). "Manipulation of cell surface macromolecules by flaviviruses". Advances in Virus Research. 59: 229–74. doi:10.1016/S0065-3527(03)59007-8. PMID 14696331.
- ↑ Owen J, Punt J, Stranford S, Jones P (2009). Immunology (7th ed.). New York: W.H. Freeman and Company. p. 423. ISBN 978-14641-3784-6.
- 1 2 3 4 Fridman WH (Sep 1991). "Fc receptors and immunoglobulin binding factors". FASEB Journal. 5 (12): 2684–90. PMID 1916092.
- ↑ Indik ZK, Park JG, Hunter S, Schreiber AD (Dec 1995). "The molecular dissection of Fc gamma receptor mediated phagocytosis". Blood. 86 (12): 4389–99. PMID 8541526.
- ↑ Harrison PT, Davis W, Norman JC, Hockaday AR, Allen JM (Sep 1994). "Binding of monomeric immunoglobulin G triggers Fc gamma RI-mediated endocytosis". The Journal of Biological Chemistry. 269 (39): 24396–402. PMID 7929100.
- 1 2 3 4 Maverakis E, Kim K, Shimoda M, Gershwin ME, Patel F, Wilken R, Raychaudhuri S, Ruhaak LR, Lebrilla CB (Feb 2015). "Glycans in the immune system and The Altered Glycan Theory of Autoimmunity: a critical review". Journal of Autoimmunity. 57 (6): 1–13. doi:10.1016/j.jaut.2014.12.002. PMC 4340844
 . PMID 25578468.
. PMID 25578468. - ↑ Zhu X, Meng G, Dickinson BL, Li X, Mizoguchi E, Miao L, Wang Y, Robert C, Wu B, Smith PD, Lencer WI, Blumberg RS (Mar 2001). "MHC class I-related neonatal Fc receptor for IgG is functionally expressed in monocytes, intestinal macrophages, and dendritic cells". Journal of Immunology. 166 (5): 3266–76. doi:10.4049/jimmunol.166.5.3266. PMC 2827247
 . PMID 11207281.
. PMID 11207281. - ↑ Firan M, Bawdon R, Radu C, Ober RJ, Eaken D, Antohe F, Ghetie V, Ward ES (Aug 2001). "The MHC class I-related receptor, FcRn, plays an essential role in the maternofetal transfer of gamma-globulin in humans". International Immunology. 13 (8): 993–1002. doi:10.1093/intimm/13.8.993. PMID 11470769.
- ↑ Simister NE, Jacobowitz Israel E, Ahouse JC, Story CM (May 1997). "New functions of the MHC class I-related Fc receptor, FcRn". Biochemical Society Transactions. 25 (2): 481–6. doi:10.1042/bst0250481. PMID 9191140.
- 1 2 3 Otten MA, van Egmond M (Mar 2004). "The Fc receptor for IgA (FcalphaRI, CD89)". Immunology Letters. 92 (1-2): 23–31. doi:10.1016/j.imlet.2003.11.018. PMID 15081523.
- ↑ Shibuya A, Honda S (Dec 2006). "Molecular and functional characteristics of the Fcalpha/muR, a novel Fc receptor for IgM and IgA". Springer Seminars in Immunopathology. 28 (4): 377–82. doi:10.1007/s00281-006-0050-3. PMID 17061088.
- ↑ Cho Y, Usui K, Honda S, Tahara-Hanaoka S, Shibuya K, Shibuya A (Jun 2006). "Molecular characteristics of IgA and IgM Fc binding to the Fcalpha/muR". Biochemical and Biophysical Research Communications. 345 (1): 474–8. doi:10.1016/j.bbrc.2006.04.084. PMID 16681999.
- ↑ Ochiai K, Wang B, Rieger A, Kilgus O, Maurer D, Födinger D, Kinet JP, Stingl G, Tomioka H (1994). "A review on Fc epsilon RI on human epidermal Langerhans cells". International Archives of Allergy and Immunology. 104 Suppl 1 (1): 63–4. doi:10.1159/000236756. PMID 8156009.
- ↑ Prussin C, Metcalfe DD (Feb 2006). "5. IgE, mast cells, basophils, and eosinophils". The Journal of Allergy and Clinical Immunology. 117 (2 Suppl Mini-Primer): S450–6. doi:10.1016/j.jaci.2005.11.016. PMID 16455345.
- ↑ von Bubnoff D, Novak N, Kraft S, Bieber T (Mar 2003). "The central role of FcepsilonRI in allergy". Clinical and Experimental Dermatology. 28 (2): 184–7. doi:10.1046/j.1365-2230.2003.01209.x. PMID 12653710.
- ↑ Kikutani H, Yokota A, Uchibayashi N, Yukawa K, Tanaka T, Sugiyama K, Barsumian EL, Suemura M, Kishimoto T (1989). "Structure and function of Fc epsilon receptor II (Fc epsilon RII/CD23): a point of contact between the effector phase of allergy and B cell differentiation". Ciba Foundation Symposium. 147: 23–31; discussion 31–5. PMID 2695308.
- ↑ Selvaraj P, Fifadara N, Nagarajan S, Cimino A, Wang G (2004). "Functional regulation of human neutrophil Fc gamma receptors". Immunologic Research. 29 (1-3): 219–30. doi:10.1385/IR:29:1-3:219. PMID 15181284.
- ↑ Sulica A, Chambers WH, Manciulea M, Metes D, Corey S, Rabinowich H, Whiteside TL, Herberman RB (1995). "Divergent signal transduction pathways and effects on natural killer cell functions induced by interaction of Fc receptors with physiologic ligands or antireceptor antibodies". Natural Immunity. 14 (3): 123–33. PMID 8832896.
- ↑ Sarfati M, Fournier S, Wu CY, Delespesse G (1992). "Expression, regulation and function of human Fc epsilon RII (CD23) antigen". Immunologic Research. 11 (3-4): 260–72. doi:10.1007/BF02919132. PMID 1287120.
- ↑ Raghavan M, Bjorkman PJ (1996). "Fc receptors and their interactions with immunoglobulins". Annual Review of Cell and Developmental Biology. 12: 181–220. doi:10.1146/annurev.cellbio.12.1.181. PMID 8970726.
- 1 2 Swanson JA, Hoppe AD (Dec 2004). "The coordination of signaling during Fc receptor-mediated phagocytosis". Journal of Leukocyte Biology. 76 (6): 1093–103. doi:10.1189/jlb.0804439. PMID 15466916.
- ↑ Sun PD (2003). "Structure and function of natural-killer-cell receptors". Immunologic Research. 27 (2-3): 539–48. doi:10.1385/IR:27:2-3:539. PMID 12857997.
- ↑ Pan L, Pei P (Apr 2003). "Signaling transduction by IgG receptors". Chinese Medical Journal. 116 (4): 487–94. PMID 12875708.
- ↑ Huang ZY, Hunter S, Kim MK, Indik ZK, Schreiber AD (Jun 2003). "The effect of phosphatases SHP-1 and SHIP-1 on signaling by the ITIM- and ITAM-containing Fcgamma receptors FcgammaRIIB and FcgammaRIIA". Journal of Leukocyte Biology. 73 (6): 823–9. doi:10.1189/jlb.0902454. PMID 12773515.
- ↑ Cambier JC (Jun 1997). "Inhibitory receptors abound?". Proceedings of the National Academy of Sciences of the United States of America. 94 (12): 5993–5. doi:10.1073/pnas.94.12.5993. PMID 9177155.
- ↑ Takai T, Ono M, Hikida M, Ohmori H, Ravetch JV (Jan 1996). "Augmented humoral and anaphylactic responses in Fc gamma RII-deficient mice". Nature. 379 (6563): 346–9. doi:10.1038/379346a0. PMID 8552190.
- ↑ Coggeshall KM (Jun 1998). "Inhibitory signaling by B cell Fc gamma RIIb". Current Opinion in Immunology. 10 (3): 306–12. doi:10.1016/s0952-7915(98)80169-6. PMID 9638367.
- ↑ Joshi T, Butchar JP, Tridandapani S (Oct 2006). "Fcgamma receptor signaling in phagocytes". International Journal of Hematology. 84 (3): 210–6. doi:10.1532/IJH97.06140. PMID 17050193.
- ↑ Trinchieri G, Valiante N (1993). "Receptors for the Fc fragment of IgG on natural killer cells". Natural Immunity. 12 (4-5): 218–34. PMID 8257828.
- ↑ Sulica A, Galatiuc C, Manciulea M, Bancu AC, DeLeo A, Whiteside TL, Herberman RB (Apr 1993). "Regulation of human natural cytotoxicity by IgG. IV. Association between binding of monomeric IgG to the Fc receptors on large granular lymphocytes and inhibition of natural killer (NK) cell activity". Cellular Immunology. 147 (2): 397–410. doi:10.1006/cimm.1993.1079. PMID 8453679.
- ↑ Yamasaki S, Saito T (2005). "Regulation of mast cell activation through FcepsilonRI". Chemical Immunology and Allergy. 87: 22–31. doi:10.1159/000087568. PMID 16107760.
- ↑ Wakahara S, Fujii Y, Nakao T, Tsuritani K, Hara T, Saito H, Ra C (Nov 2001). "Gene expression profiles for Fc epsilon RI, cytokines and chemokines upon Fc epsilon RI activation in human cultured mast cells derived from peripheral blood". Cytokine. 16 (4): 143–52. doi:10.1006/cyto.2001.0958. PMID 11792124.
- ↑ Metcalfe DD, Baram D, Mekori YA (Oct 1997). "Mast cells". Physiological Reviews. 77 (4): 1033–79. PMID 9354811.
- ↑ David JR, Butterworth AE, Vadas MA (Sep 1980). "Mechanism of the interaction mediating killing of Schistosoma mansoni by human eosinophils". The American Journal of Tropical Medicine and Hygiene. 29 (5): 842–8. PMID 7435788.
- ↑ Capron M, Soussi Gounni A, Morita M, Truong MJ, Prin L, Kinet JP, Capron A (1995). "Eosinophils: from low- to high-affinity immunoglobulin E receptors". Allergy. 50 (25 Suppl): 20–3. doi:10.1111/j.1398-9995.1995.tb04270.x. PMID 7677229.
- ↑ Gounni AS, Lamkhioued B, Ochiai K, Tanaka Y, Delaporte E, Capron A, Kinet JP, Capron M (Jan 1994). "High-affinity IgE receptor on eosinophils is involved in defence against parasites". Nature. 367 (6459): 183–6. doi:10.1038/367183a0. PMID 8114916.
Further reading
- Laneway, Charles A.; Travers, Paul; Waldport, Mark; Shlomchik, Mark J (2001). "Chapter 9. The Humoral Immune Response". Immunobiology: The Immune System in Health and Disease (5th ed.). New York: Garland. ISBN 978-0-8153-3642-6.
- Abbas AK, Lichtman AH, Pillai S (2012). "Chapter 12: Effector Mechanisms of Humoral Immunity". Cellular and molecular immunology (7th ed.). Philadelphia: Elsevier/Saunders. ISBN 978-1-4377-1528-6.
- Gerber JS, Mosser DM (Feb 2001). "Stimulatory and inhibitory signals originating from the macrophage Fcgamma receptors". Microbes and Infection / Institut Pasteur. 3 (2): 131–9. doi:10.1016/s1286-4579(00)01360-5. PMID 11251299.
- Maverakis E, Kim K, Shimoda M, Gershwin ME, Patel F, Wilken R, Raychaudhuri S, Ruhaak LR, Lebrilla CB (Feb 2015). "Glycans in the immune system and The Altered Glycan Theory of Autoimmunity: a critical review". Journal of Autoimmunity. 57: 1–13. doi:10.1016/j.jaut.2014.12.002. PMC 4340844
 . PMID 25578468.
. PMID 25578468.
External links
- Fc Receptor at the US National Library of Medicine Medical Subject Headings (MeSH)